
Whole Plasmid Sequencing is offered through our parent company, Eurofins Genomics. All links below will redirect to www.eurofinsgenomics.com. You will need to create a new account with Eurofins Genomics to place orders.
Fast, accurate, and affordable long-read sequencing
Confirm full plasmids faster, more accurately, and more affordably than ever before with Whole Plasmid Sequencing. NGS generation 3 technology has made it possible to quickly sequence whole plasmids in a fraction of the time and without the hassle of having to design and synthesize primers. The typical turnaround time for whole plasmid sequencing is 1 business day measured from when the samples are received. There is no minimum requirement for the number of samples.

Same Day Results
Results are Delivered the Same Day Samples are received

$15 Per Plasmid
Sequencing Verification for a fraction of the price
Full Length Plasmids
Get the complete picture of your plasmid from 5kb – 30kb
SAME GREAT SEQUENCING / WITHOUT THE PRIMERS
Sample Submission Guidelines
| Sample type | Size Categor | Length | Concentration | Min volume | Price per sample |
|---|---|---|---|---|---|
| Plasmid | Regular | 2.5 - 25 kb | 30 ng/uL | ≥10 uL | $15 |
| Large | 25 - 125 kb | 50 ng/uL | ≥20 uL | $30 | |
| XL | 125 - 300 kb | 50 ng/uL | ≥40 uL | $60 |
*NOTE: this order button will redirect to the Eurofins Genomics website
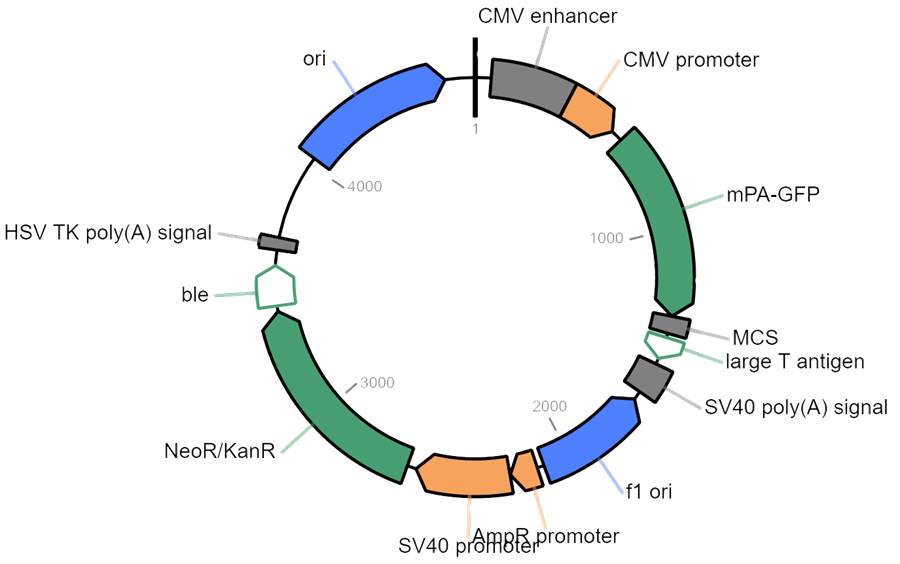
How do I ship samples?
1. Dropbox – we have a nationwide network of dropboxes. If you want a box for your lab, let us know! Email GenomicsSupport@eurofins.com to request a box.
2. Free Digital Shipping Labels – digital shipping labels are provided free for a vast majority of sequencing orders. You can select the option for digital shipping label during checkout.
3. Ship the samples using your normal carrier – If using your own carrier, we highly recommend shipping overnight/next-day delivery to ensure the samples do not degrade in transit.
How do I get results?
Results are available to download from the order history page. You will be emailed when the results are ready as well. Currently, 95% of results are delivered within 1 business day from receiving the sample.
THE EASIEST POSSIBLE WAY TO / SEQUENCE

No Primers Needed
Unlike short-read sequencing, whole plasmid sequencing does not require any primers.

No library prep needed
Unlike conventional next generation sequencing methods, no library prep is required.

No Shipping Cost
You can use our existing network of dropboxes to submit samples or print a free digital shipping label at checkout. If you would like a dropbox for your lab, let us know! Email GenomicsSupport@eurofins.com to request a box.
Accuracy
Oxford Nanopore’s reports the raw read accuracy is 98.3% and the consensus accuracy for SNPs is 99.6% with 50X coverage.
BENEFITS / AND ADVANTAGES
Benefits
– Long read recovery of several kilobases
– Great for GC rich or repetitive DNA
– Detect E. coli and genomic DNA carryover
– Faster TAT compared to traditional primer walking and plasmid verification techniques
– Lower cost than traditional NGS methods
– Verification of the entire vector sequence
– Scalable
– Available from one sample
– 5 – 4 million reads per Flow Cell (dependent on DNA quality and length)
Applications
Whole plasmid sequencing using NGS Gen 3 technology is a powerful and accurate method for characterizing and analyzing plasmids, which makes it well suited for a wide range of applications.
– Plasmid Verification
– Resequencing of whole genomes
– Assembly of genomes
– Identification of taxonomic background
– Metagenomic analysis of long reads
– Much more.
Easy to Submit
– No primers required! It could not be easier
– Requires less DNA than short-read sequencing
– Super simple order page
– Turnaround is typically 1 business day, measured from when samples arrive.
– Nationwide network of dropboxes
– No dropbox near you? No problem. Eurofins Genomics is the only company that offers free digital shipping labels.
FAQ / DETAILS
Currently >90% of results are delivered by 5pm ET the same day the samples are received.
Any number of samples is welcome. There are no minimums or limitations.
Dropboxes are available in many locations. Also, Eurofins is the only sequencing company that offers free digital shipping labels on >95% of orders. Take a look at our sample submission page for details.
The equipment vendor reports accuracy of 99.3%. The research community has reported accuracy between 93-98%.
There are pros and cons of every sequencing method. Sanger sequencing and Oxford Nanopore sequencing (ONT) are both methods for determining the sequence of nucleotides in a piece of DNA. Typically Sanger is consider the most accurate method for short-read sequencing and NGS is better for long-read sequencing.
Overall, the choice between Sanger sequencing and ONT will depend on the specific needs of the application. Both methods have their strengths and limitations, and the appropriate method will depend on factors such as the length and quality of the DNA sample, the desired level of accuracy, and the cost and availability of the necessary equipment.
Pros | Cons | |
Short read sequencing (Sanger) |
|
|
Long read sequencing (Whole Plasmid) |
|
|
| Sample type | Size Category | Length | Concentration | Min volume | Price per sample |
| Plasmid | Regular | 2.5 – 25 kb | 30 ng/uL | ≥10 uL | $15 |
| Large | 25 – 125 kb | 50 ng/uL | ≥20 uL | $30 | |
| XL | 125 – 300 kb | 50 ng/uL | ≥40 uL | $60 |
Absolutely. Using our free barcode labels is not required, but always an option.
Absolutely, yes.
You can send it and we can sequence it, but we cannot predict or promise the analysis outcome. The customer would take on the risk of these orders.
We will be rolling out the new order page across portal sites, marketplaces, and punchouts very soon. At the moment, please submit orders on the main www.eurofinsgenomics.com website.
It really depends on the sample quality. We cannot guarantee the level of coverage at this time. A good sample submitted properly will typically yields hundreds or even thousands of sequencing reads. Consensus coverage depends on how many reads are full-length plasmids and how many, if any, degraded.
- .fasta file (for consensus data): we will provide a clean, complete consensus sequence for each plasmid.
- .gbk GenBank file (for consensus data): a pLannotate map in the GenBank file format.
- (OPTIONAL) .fastq file – raw data on reads. Email support to request fastq files.
- Histogram file: the hisogram file provides a visual representation of the plasmid and raw read data for deeper insight into your samples (image).
- .html pLannotate map (for consensus data): a plasmid map for each sample.
The probability of failure is low when using WPS, although it can still happen. If you wish to resequence a failed sample, contact Genomics Support. Unfortunately, we must charge for failed samples since it requires more time and energy than a normal run on the machine. If the sample fails a second time, all data points to a problem with the sample and we cannot resequence. In that scenario, we advise the customer to either send a new sample or troubleshoot the sample from their end.
We will be adding this service soon but not yet.
NOTE: this order button will redirect to the Eurofins Genomics website
