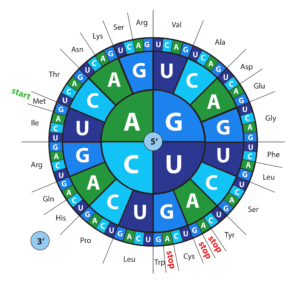
Codon Optimization
Eurofins Genomics Blue Heron has developed a proprietary algorithm for protein expression optimization. Based on codon usage and comprehensive knowledge of optimal RNA structure for protein translation, we can confidently deliver results in the desired organism.
How does Eurofins Genomics Blue Heron's proprietary algorithm "Codon" optimize my sequence?
First, Eurofins Genomics Blue Heron’s proprietary sequence optimization algorithm removes rare codons from the customer designated codon utilization table of a selected species. Then the remaining codon utilization frequencies are normalized. Next, the amino acid sequence is reverse translated into a DNA sequence by sampling frequently used codons from the normalized codon utilization table while including or excluding any DNA sequences (including restriction sites) specified by the customer. The resulting output is a DNA sequence, with specified included and excluded sequences, that is codon optimized for the desired species. Based on feedback from our customers, we recommend codon optimization when you are planning to use a mammalian host for expression of the protein.
How does the proprietary algorithm "Expression" optimize my sequence?
First the algorithm codon optimizes the amino acid sequence as described above. Then, the algorithm continues to iteratively sample codons based on the probability distribution of the normalized codon utilization table to find a low free energy solution. The resulting output is a DNA sequence with specified included or excluded DNA sequences and reduced secondary structure of the corresponding mRNA. We recommend expression optimization for heterologous expression or when secondary structure is likely to impede translation.
Advantages
- Reliable – a proven algorithm: successfully used by many researchers.
- Flexible – exclude unwanted restriction sites and special sequences, use a customer’s codon table, and add flanking sequences.
- Free – no charge for registered users.
- Easy to use.
Procedure
- Login, or create a new account.
- Submit a protein sequence.
- Select your model organism or your own codon table.
- Check any restriction sites to be avoided (optional).
- Enter any special sequences to be avoided (optional).
- Add flanking sequences (optional).
- An optimized sequence will be emailed and posted on your secure account.
How can I calculate the free energy of my native or base DNA sequence?
Eurofins Genomics Blue Heron uses the Vienna RNA Package to calculate the free energy of the RNA sequence that corresponds to your DNA sequence. This free energy value is used during expression optimization as a measure of the RNA secondary structure. You can download a copy of this software from the Vienna RNA Package Website. If you wish to calculate the free energy of your sequence prior to our Expression Optimization, you can install and use the RNAfold program from the Vienna RNA package. Please see the Vienna RNA Website given above for the full RNAfold documentation.